Write-Audit-Publish for Data Lakes in Pure Python (no JVM)
Previously published in Towards Data Science on 04/11/2024.

Introduction
In this blog post we provide a no-nonsense, reference implementation for Write-Audit-Publish (WAP) patterns on a data lake, using Apache Iceberg as an open table format, and Project Nessie as a data catalog supporting git-like semantics.
We chose Nessie because its branching capabilities provide a good abstraction to implement a WAP design. Most importantly, we chose to build on PyIceberg to eliminate the need for the JVM in terms of developer experience. In fact, to run the entire project, including the integrated applications we will only need Python and AWS.
While Nessie is technically built in Java, the data catalog is run as a container by AWS Lightsail in this project, we are going to interact with it only through its endpoint. Consequently, we can express the entire WAP logic, including the queries downstream, in Python only!
Because PyIceberg is fairly new, a bunch of things are actually not supported out of the box. In particular, writing is still in early days, and branching Iceberg tables is still not supported. So what you'll find here is the result of some original work we did ourselves to make branching Iceberg tables in Nessie possible directly from Python.
So all this happened, more or less.
What on earth is WAP?
Back in 2017, Michelle Winters from Netflix talked about a design pattern called Write-Audit-Publish (WAP) in data. Essentially, WAP is a functional design aimed at making data quality checks easy to implement before the data becomes available to downstream consumers.
For instance, a typical use case is data quality at ingestion. The flow will look like creating a staging environment and running quality tests on freshly ingested data, before making that data available to any downstream application.
As the name suggests, there are essentially three phases:
- Write. Put the data in a location that is not accessible to consumers downstream (e.g., a staging environment or a branch).
- Audit. Transform and test the data to make sure it meets the specifications (e.g., check whether the schema abruptly changed, or whether there are unexpected values, such as NULLs).
- Publish. Put the data in the place where consumers can read it from (e.g., the production data lake).
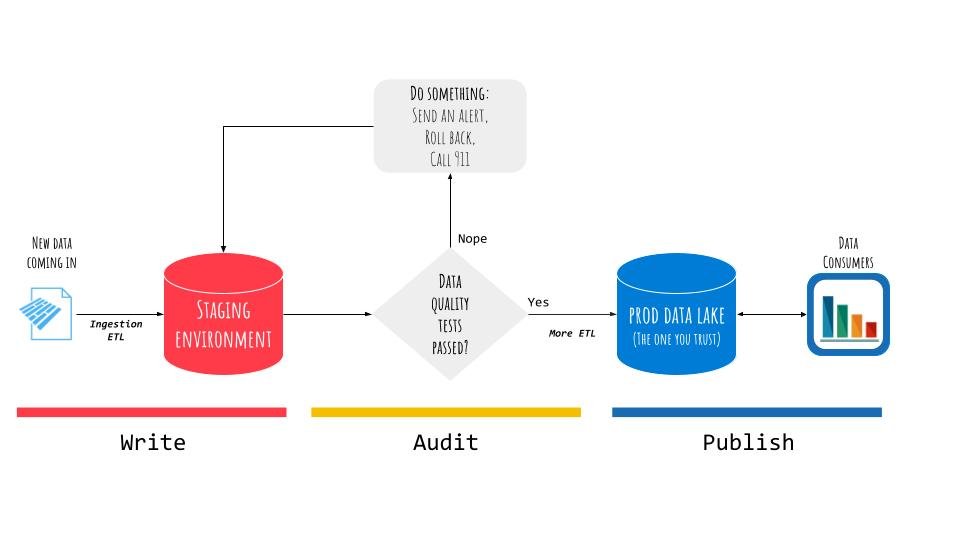
This is only one example of the possible applications of WAP patterns. It is easy to see how it can be applied at different stages of the data life-cycle, from ETL and data ingestion, to complex data pipelines supporting analytics and ML applications.
Despite being so useful, WAP is still not very widespread, and only recently companies have started thinking about it more systematically. The rise of open table formats and projects like Nessie and LakeFS is accelerating the process, but it still a bit avant garde.
In any case, it is a very good way of thinking about data and it is extremely useful in taming some of the most widespread problems keeping engineers up at night. So let’s see how we can implement it.
WAP on a data lake in Python
We are not going to have a theoretical discussion about WAP nor will we provide an exhaustive survey of the different ways to implement it (Alex Merced from Dremio and Einat Orr from LakeFs are already doing a phenomenal job at that). Instead, we will provide a reference implementation for WAP on a data lake.
👉 So buckle up, clone the Repo, and give it a spin!
📌 For more details, please refer to the README of the project.
Architecture and workflow
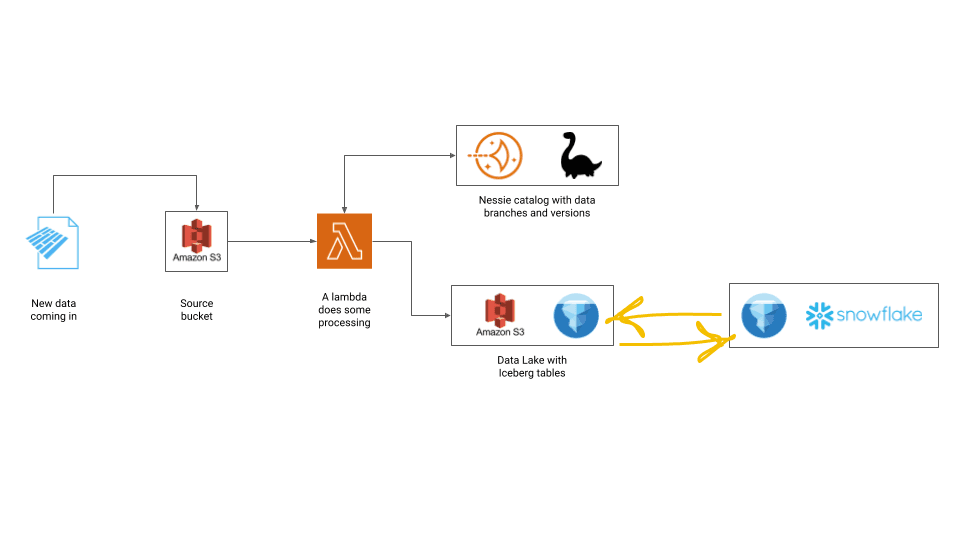
The idea here is to simulate an ingestion workflow and implement a WAP pattern by branching the data lake and running a data quality test before deciding whether to put the data into the final table into the data lake.
We use Nessie branching capabilities to get our sandboxed environment where data cannot be read by downstream consumers and AWS Lambda to run the WAP logic.
Essentially, each time a new parquet file is uploaded, a Lambda will go up, create a branch in the data catalog and append the data into an Iceberg table. Then, a simple a simple data quality test is performed with PyIceberg to check whether a certain column in the table contains some NULL values.
If the answer is yes, the data quality test fails. The new branch will not be merged into the main branch of the data catalog, making the data impossible to be read in the main branch of data lake. Instead, an alert message is going to be sent to Slack.
If the answer is no, and the data does not contain any NULLs, the data quality test is passed. The new branch will thus be merged into the main branch of the data catalog and the data will be appended in the Iceberg table in the data lake for other processes to read.
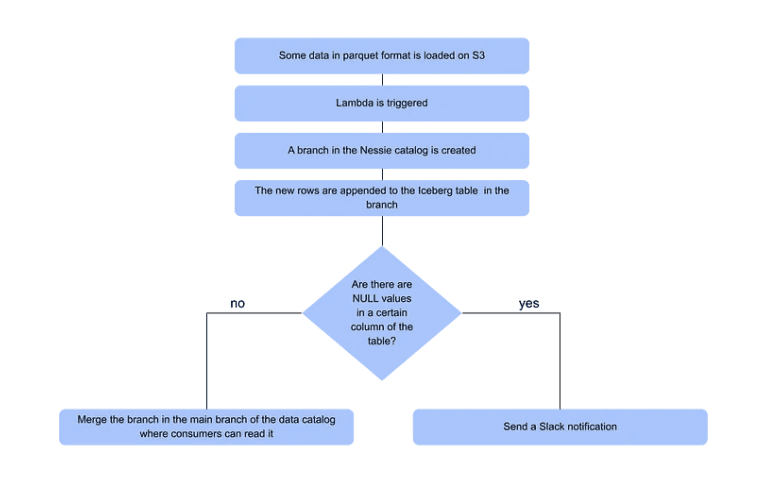
All data is completely synthetic and is generated automatically by simply running the project. Of course, we provide the possibility of choosing whether to generate data that complies with the data quality specifications or to generate data that include some NULL values.
To implement the whole end-to-end flow, we are going to use the following components:
- Storage: AWS S3
- Open table format: Apache Iceberg
- Data catalog: Project Nessie
- Code implementation: PyIceberg, PyNessie
- Serverless runtime: Lambda
- Virtual private server: Lightsail
- Alerting system: Slack
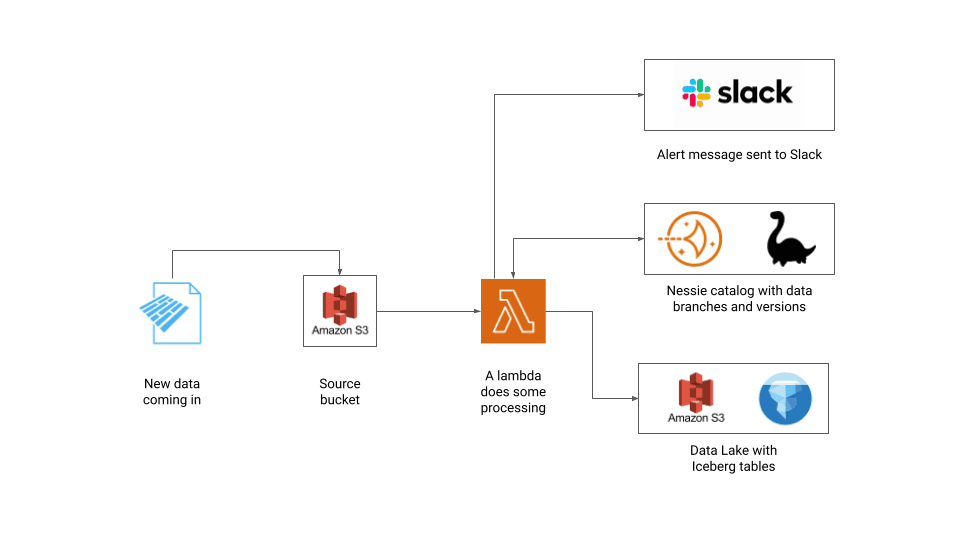
This project is pretty self-contained and comes with scripts to set up the entire infrastructure, so it requires only introductory-level familiarity with AWS and Python.
It’s also not intended to be a production-ready solution, but rather a reference implementation, a starting point for more complex scenarios: the code is verbose and heavily commented, making it easy to modify and extend the basic concepts to better suit anyone’s use cases.
Visualize
To visualize the results of the data quality test, we provide a very simple Streamlit app that can be used to see what happens when some new data is uploaded to first location on S3 — the one that is not available to downstream consumers.
We can use the app to check how many rows are in the table across the different branches, and for the branches other than main, it is easy to see in what column the data quality test failed and in how many rows.
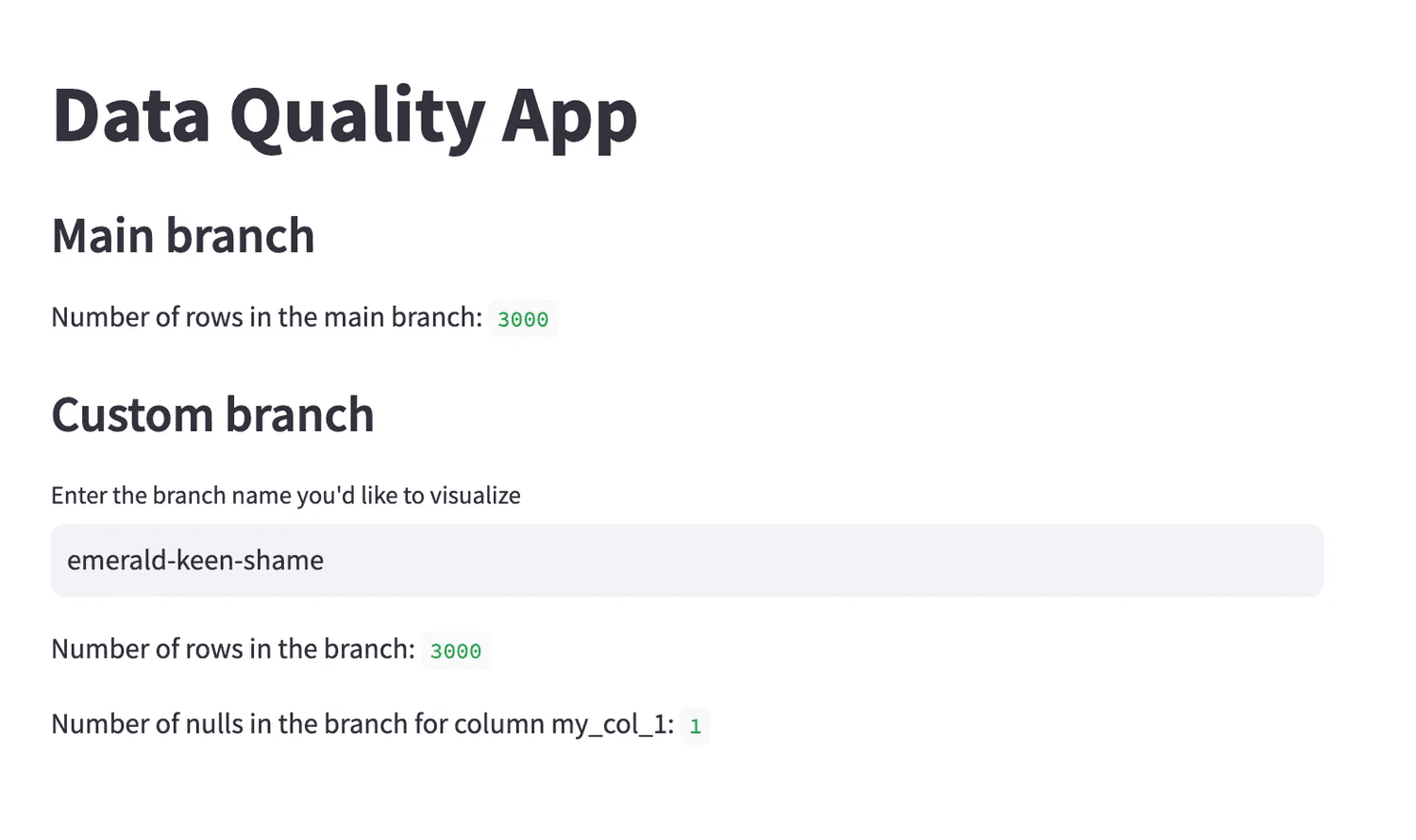
From the lake to the Lakehouse
Once we have a WAP flow based on Iceberg, we can leverage it to implement a composable design for our downstream consumers. In our repo we provide instructions for a Snowflake integration as a way to explore this architectural possibility.
This is one of the main tenet of the Lakehouse architecture, conceived to be more flexible than modern data warehouses and more usable than traditional data lakes.
On the one hand, the Lakehouse hinges on leveraging object store to eliminate data redundancy and at the same time lower storage cost. On the other, it is supposed to provide more flexibility in choosing different compute engines for different purposes.
All this sounds very interesting in theory, but it also sounds very complicated to engineer at scale. Even a simple integration between Snowflake and an S3 bucket as an external volume is frankly pretty tedious.
And in fact, we cannot stress this enough, moving to a full Lakehouse architecture is a lot of work. Like a lot!
Having said that, even a journey of a thousand miles begins with a single step, so why don’t we start by reaching out the lowest hanging fruits with simple but very tangible practical consequences?
The example in the repo showcases one of these simple use case: WAP and data quality tests. The WAP pattern here is a chance to move the computation required for data quality tests (and possibly for some ingestion ETL) outside the data warehouse, while still maintaining the possibility of taking advantage of Snowflake for more high value analyitcs workloads on certified artifacts. We hope that this post can help developers to build their own proof of concepts and use the
Conclusions
The reference implementation here proposed has several advantages:
Tables are better than files
Data lakes are historically hard to develop against, since the data abstractions are very different from those typically adopted in good old databases. Big Data frameworks like Spark first provided the capabilities to process large amounts of raw data stored as files in different formats (e.g. parquet, csv, etc), but people often do not think in terms of files: they think it terms of tables.
We use an open table format for this reason. Iceberg turns the main data lake abstraction into tables rather than files which makes things considerably more intuitive. We can now use SQL query engines natively to explore the data and we can count on Iceberg to take care of providing correct schema evolution.
Interoperability is good for ya
Iceberg also allows for greater interoperability from an architectural point of view. One of the main benefits of using open table formats is that data can be kept in object store while high-performance SQL engines (Spark, Trino, Dremio) and Warehouses (Snowflake, Redshift) can be used to query it. The fact that Iceberg is supported by the majority of computational engines out there has profound consequences for the way we can architect our data platform.
As described above, our suggested integration with Snowflake is meant to show that one can deliberately move the computation needed for the ingestion ETL and the data quality tests outside of the Warehouse, and keep the the latter for large scale analytics jobs and last mile querying that require high performance. At scale, this idea can translate into significantly lower costs.
Branches are useful abstractions
WAP pattern requires a way to write data in a location where consumers cannot accidentally read it. Branching semantics naturally provides a way to implement this, which is why we use Nessie to leverage branching semantics at the data catalog level. Nessie builds on Iceberg and on its time travel and table branching functionalities. A lot of the work done in our repo is to make Nessie work directly with Python. The result is that one can interact with the Nessie catalog and write Iceberg tables in different branches of the data catalog without a JVM based process to write.
Simpler developer experience
Finally, making the end-to-end experience completely Python-based simplifies remarkably the set up fo the system and the interaction with it. Any other system we are aware of would require a JVM or an additional hosted service to write back into Iceberg tables into different branches, while in this implementation the entire WAP logic can run inside one single lambda function.
There is nothing inherently wrong with the JVM. It is a fundamental component of many Big Data frameworks, providing a common API to work with platform-specific resources, while ensuring security and correctness. However, the JVM takes a toll from a developer experience perspective. Anybody who worked with Spark knows that JVM-based systems tend to be finicky and fail with mysterious errors. For many people who work in data and consider Python as their lingua franca the advantage of the JVM is paid in the coin of usability.
We hope more people are excited about composable designs like we are, we hope open standards like Iceberg and Arrow will become the norm, but most of all we hope this is useful.
So it goes.







